Research and publish the best content.
Get Started for FREE
Sign up with Facebook Sign up with X
I don't have a Facebook or a X account
Already have an account: Login
 Your new post is loading... Your new post is loading...
 Your new post is loading... Your new post is loading...
|
|










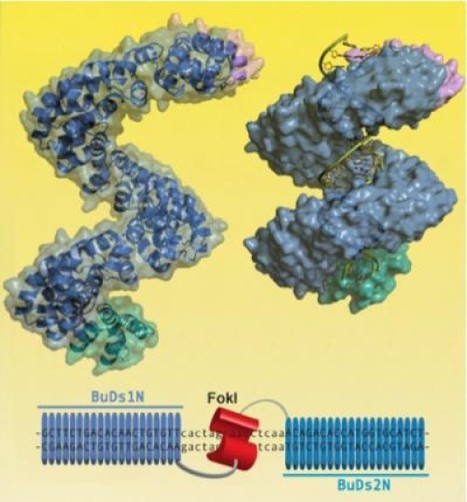




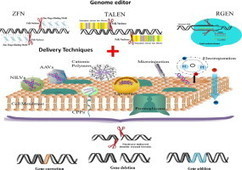
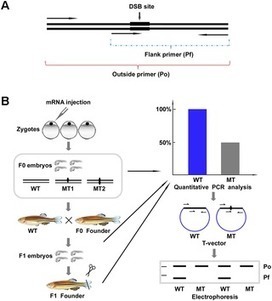


The authors discuss experimental considerations, limitations and critical aspects which will guide the investigator for successful implementation of the genome editing technology in human PSCs using designer nucleases.