CRISPR knockout screens outperform shRNA and CRISPR-interference screens in a side-by-side comparison.
Get Started for FREE
Sign up with Facebook Sign up with X
I don't have a Facebook or a X account
 Your new post is loading... Your new post is loading...
 Your new post is loading... Your new post is loading...
|

Martina Hrabinová's curator insight,
March 31, 2016 5:14 AM
A new way to avoid genotoxicity? Several types of CRISPR–Cas adaptive immune system have been described, each with a unique mechanism and protein composition. DNA cleavage by type III-A CRISPR–Cas systems relies on a complex known as Cas10–Csm and requires transcription of the target DNA. Type III-A CRISPR–Cas systems are also capable of degrading RNA, although why they should have an RNase function is not known. Jiang et al. now show that some phage targets can delay the transcription-dependent DNA cleavage mechanism of type III-A CRISPR–Cas systems but that CRISPR RNases prevent the replication of these phages. |
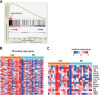















High-throughput genetic screens have become essential tools for studying a wide variety of biological processes. Here the scientists experimentally compare systems based on clustered regularly interspaced short palindromic repeat (CRISPR)/CRISPR-associated protein 9 (Cas9) or its transcriptionally repressive variant, CRISPR-interference (CRISPRi), with a traditional short hairpin RNA (shRNA)-based system for performing lethality screens. They find that the CRISPR technology performed best, with low noise, minimal off-target effects and consistent activity across reagents.