Enhancing CRISPR-mediated site-specific transgene insertion efficiency by homology-directed repair (HDR) using high concentrations of double-stranded DNA (dsDNA) with Cas9 target sequences (CTSs) can be toxic to primary cells. Here, we develop single-stranded DNA (ssDNA) HDR templates (HDRTs) incorporating CTSs with reduced toxicity that boost knock-in efficiency and yield by an average of around two- to threefold relative to dsDNA CTSs. Using small-molecule combinations that enhance HDR, we could further increase knock-in efficiencies by an additional roughly two- to threefold on average. Our method works across a variety of target loci, knock-in constructs and primary human cell types, reaching HDR efficiencies of >80–90%. We demonstrate application of this approach for both pathogenic gene variant modeling and gene-replacement strategies for IL2RA and CTLA4 mutations associated with Mendelian disorders. Finally, we develop a good manufacturing practice (GMP)-compatible process for nonviral chimeric antigen receptor-T cell manufacturing, with knock-in efficiencies (46–62%) and yields (>1.5 × 109 modified cells) exceeding those of conventional approaches. Combinations of single-stranded DNA repair templates and small molecules markedly enhance genome editing.
Research and publish the best content.
Get Started for FREE
Sign up with Facebook Sign up with X
I don't have a Facebook or a X account
Already have an account: Login
 Your new post is loading... Your new post is loading...
 Your new post is loading... Your new post is loading...
|
|




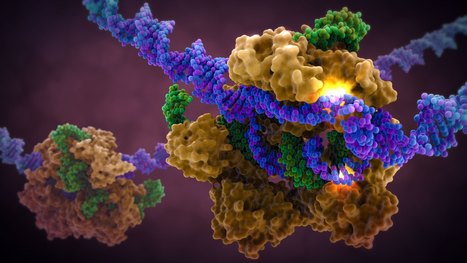





A new approach to CRISPR-Cas9 technology allows scientists to introduce particularly long DNA sequences at precise locations in cell genomes at remarkably high efficiencies without the viral delivery systems traditionally used to transport DNA into cells. The researchers found that high levels of double-stranded DNA template can be toxic to cells, resulting in low efficiency. The team knew that single-stranded DNA was less toxic to cells, even at relatively high concentrations. The researchers therefore describe in the paper published in the journal Nature Biotechnology, a method for attaching the modified Cas9 enzyme to a single-stranded template DNA, by adding just a small overhang of double-stranded DNA at the ends. The single-stranded template DNA could more than double the efficiency of gene editing compared to the old double-stranded approach. In addition, the double-stranded ends of the molecules allow the researchers to use Cas9 to enhance the delivery of non-viral vectors into cells. In the study, the researchers used the new DNA template to generate more than one billion CAR-T cells targeting multiple myeloma and showed that their approach could, for the first time, replace two genes associated with rare genetic immune diseases, IL2RA and CTLA4, in their entirety