Immunotherapy targeting amyloid-β peptide is under active clinical investigation for treatment of Alzheimer’s disease (AD). Among the hypotheses being investigated for impact on clinical outcome are the preferred epitope or conformation of amyloid-β to target for treatment, and the mechanism of action underlying immunotherapy. Bapineuzumab (humanized 3D6), a neo-epitope specific antibody recognizing amyloid-β1-5 with strong preference for an exposed Asp residue at the N-terminus of the peptide, has undergone advanced clinical testing for treatment of AD.
Here we report the crystal structure of a recombinant Fab fragment of 3D6 in complex with amyloid-β1-7 solved at 2.0 Å resolution. The N-terminus of amyloid-β is bound to 3D6 as a 310 helix. The amino-terminal Asp residue is buried deepest in the antibody binding pocket, with the Cβ atom of residue 6 visible at the entrance to the binding pocket near the surface of the antibody. We further evaluate homology model based predictions used to guide humanization of 3D6 to bapineuzumab, with actual structure of the Fab. The structure of the Fab:amyloid-β complex validates design of the humanized antibody, and confirms the amyloid-β epitope recognized by 3D6 as previously mapped by ELISA.
ConclusionsThe conformation of amyloid-β antigen recognized by 3D6 is novel and distinct from other antibodies recognizing N-terminal epitopes. Our result provides the first report demonstrating structural conservation of antigen contact residues, and conformation of antigen recognized, between the parent murine antibody and its humanized version.
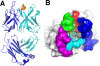
Figure 1. Overview of the structure of 3D6 fab with Aβ1-7 peptide (pdb identifier 4ONF). A) Side view showing alpha-carbon backbone traces of the molecules. Heavy chain is shown in cyan, light chain in blue and Aβ peptide in yellow/orange. B) 3D6 with Aβ1-7 view from above the molecule. Peptide is shown in stick representation with oxygens colored red, nitrogens in blue and carbons orange. Fab in surface representation colored white with the exception of CDRs contacting the peptide. CDR H1 is shown in red, H2 blue, H3 green, L1 magenta and L3 cyan. Only CDRs contacting the peptide are colored. CDRs, complementarity determining regions; pdb, protein data base.
Via Krishan Maggon


 Your new post is loading...
Your new post is loading...




open access
Crystal structure reveals conservation of amyloid-β conformation recognized by 3D6 following humanization to bapineuzumabHadar Feinberg2, José W Saldanha5, Linnea Diep1, Amita Goel1, Angela Widom3,Geertruida M Veldman6, William I Weis2, Dale Schenk4 and Guriqbal S Basi1*
*Corresponding author: Guriqbal S Basi gur1balbasi@gmail.com
Author Affiliations
1Elan Pharmaceuticals, Inc. 300 Technology Sq., Cambridge, MA 02139, USA
2Departments of Structural Biology and of Molecular & Cellular Physiology, 299 Campus Drive, Stanford University School of Medicine, Stanford, CA 94305, USA
3Pfizer, Cambridge, MA, USA
4Prothena Biosciences, Inc., 650 Gateway Blvd., San Francisco, CA 94080, USA
5National Institute for Medical Research, The Ridgeway, Mill Hill, London NW7 1AA, UK
6Abbvie Bioresearch Center, Worcester, MA, USA
For all author emails, please log on.
Alzheimer's Research & Therapy 2014, 6:31 doi:10.1186/alzrt261